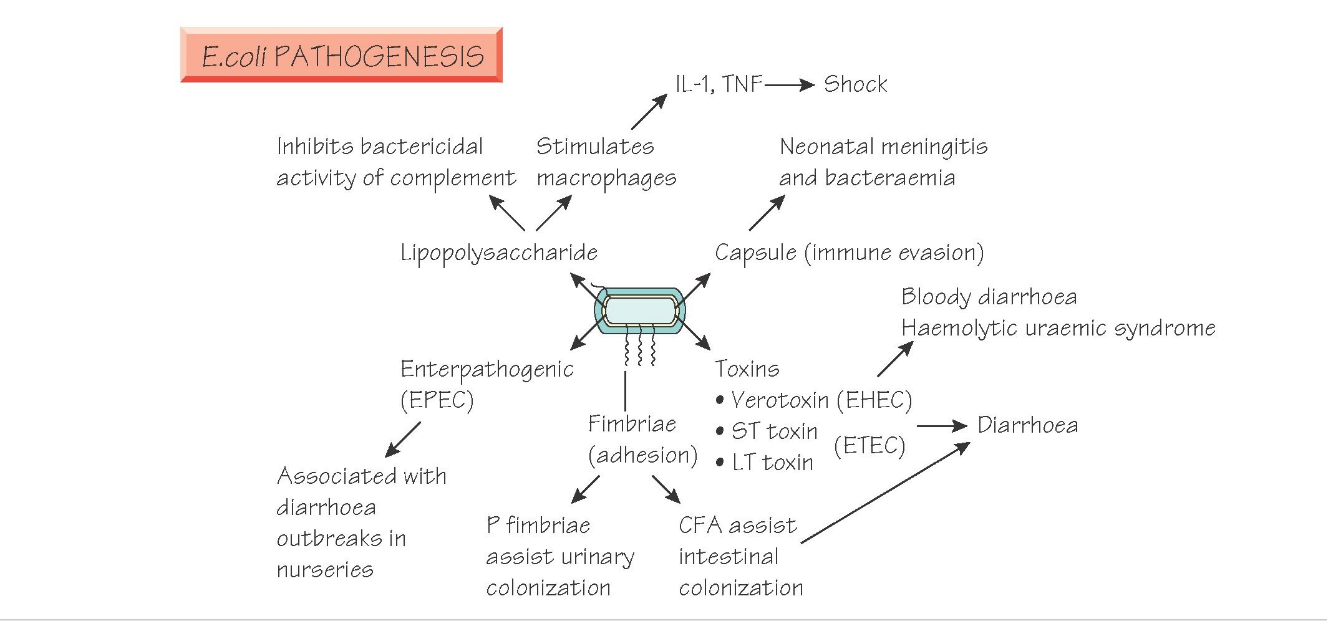
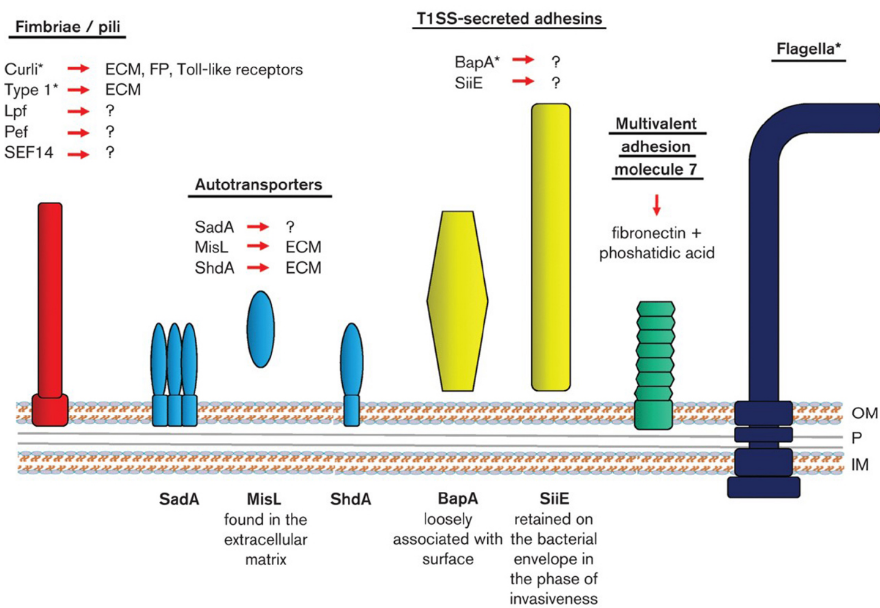
Pathogenicity of Gram-negative Pathogens.
Neil R. Baker baker.2@osu.edu Associate Professor, Ph.D., University of Maryland, 1975. Pathogenicity of gram-negative pathogens. My research is primarily concerned with secondary pathogens of the respiratory tract. Pseudomonas aeruginosa, has been of particular concern because of the high mortality rates associated with hospital-acquired infections and the high incidence in cystic